In this work the Xing Lab tackled an outstanding open question on if and how one can extract dynamical information from snapshot data.
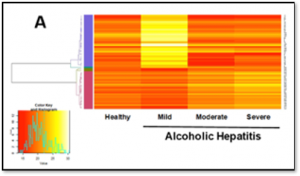
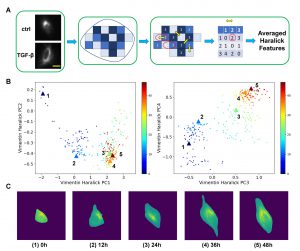
They first developed a quantitative framework that integrates standard imaging facilities and state-of-the-art computational analysis approaches to extract high-dimensional dynamical features of single live cell trajectories. The ability of being “quantitative” and “high-dimensional” is critical for addressing the question mentioned above. The framework allows one to use the same mathematical language to quantitatively describe cell phenotypic transition dynamics as one describes particle motions in physics and chemistry. This a conceptual novelty sets up a new framework of studying the biological processes from a physics perspective. They studied the epithelial-to-mesenchymal transition, and identified two parallel paths for the transition process that are concealed from snapshot data due to cell-cell heterogeneity. The work demonstrates the importance of live cell studies, and our developed framework provides such a general quantitative platform.
Wang W, Douglas D, Zhang J, Chen YJ, Cheng YY, Kumari S, Enuameh MS, Dai Y, Wallace CT, Watkins SC, Shu W, Xing J. (2020) Live cell imaging and analysis reveal cell phenotypic transition dynamics inherently missing in snapshot data. Science Advances.