De novo emergence of adaptive membrane proteins from thymine-rich genomic sequences
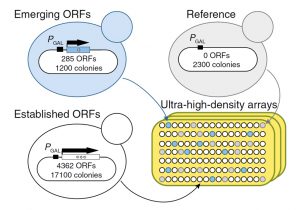
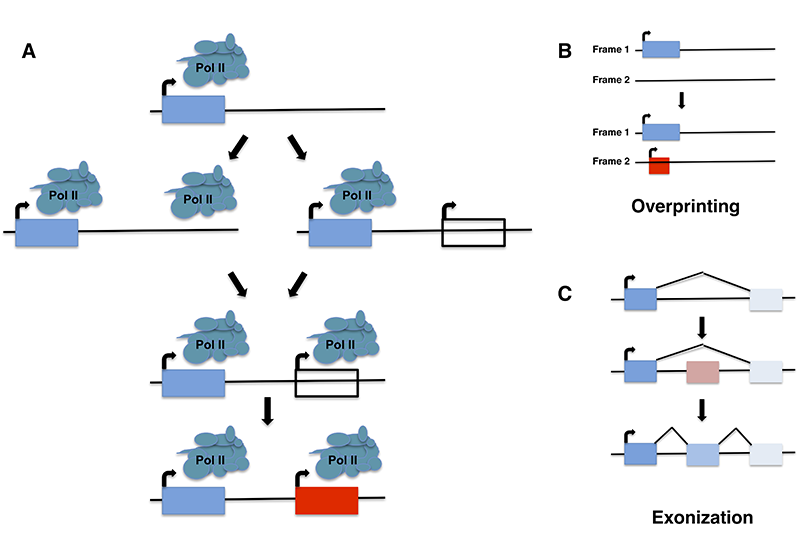
Recent evidence demonstrates that novel protein-coding genes can arise de novo from nongenic loci. This evolutionary innovation is thought to be facilitated by the pervasive translation of non-genic transcripts, which exposes a reservoir of variable polypeptides to natural selection. We find that adaptive emerging sequences tend to encode putative transmembrane domains, and that thymine-rich intergenic regions harbor a widespread potential to produce transmembrane domains. These findings, together with in-depth examination of the de novo emerging YBR196C-A locus, suggest a novel evolutionary model whereby adaptive transmembrane polypeptides emerge de novo from thymine-rich nongenic regions and subsequently accumulate changes molded by natural selection.

Vakirlis N, Acar O, Hsu B, Coelho NC, Van Oss SB, Wacholder A, Medetgul-Ernar K, Bowman II RW, Hines CP, Iannotta J, Parikh SB, McLysaght A, Camacho CJ, O’Donnell AF, Ideker T, Carvunis AR. De novo emergence of adaptive membrane proteins from thymine-rich genomic sequences. Nat Commun 11, 781 (2020). https://doi.org/10.1038/s41467-020-14500-z