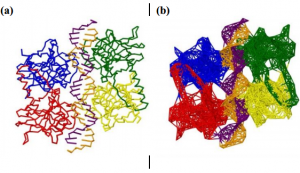
 | Eyal E, Lum G, Bahar I (2015) The Anisotropic Network Model web server at 2015 (ANM 2.0) Bioinformatics 31(9): 1487-9. |
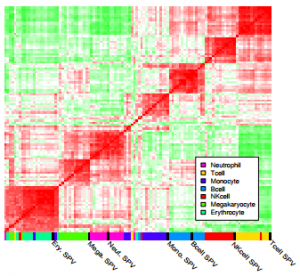 | Chikina M, Zaslavsky E, Sealfon SC (2015) CellCODE: A robust latent variable approach to differential expression analysis for heterogeneous cell populations Bioinformatics 31(10): 1584-91. |
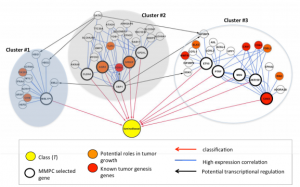 | Huang GT, Tsamardinos I, Raghu V, Kaminski N, Benos PV (2015) T-recs: stable selection of dynamically formed groups of features with application to prediction of clinical outcomes Pac Symp Biocomput 20: 431-42. |
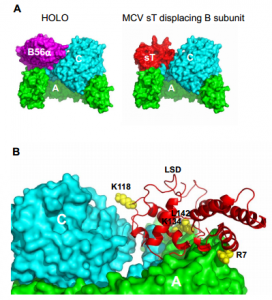 | Kwun HJ, Shuda M, Camacho CJ, Gamper AM, Thant M, Chang Y, Moore PS (2015) Restricted Protein Phosphatase 2A Targeting by Merkel Cell Polyomavirus Small T Antigen J Virol pii: JVI.00157-15 89(8): 4191-200. |
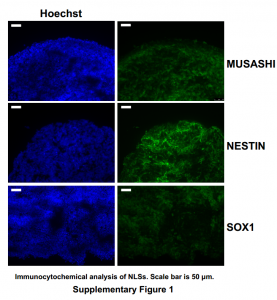 | D'Aiuto L, Zhi Y, Das DK, Wilcox MR, Johnson JW, McClain L, MacDonald ML, Di Maio R, Schurdak ME, Piazza P, Viggiano L, Sweet R, Kinchington PR, Bhattacharjee AG, Yolken R, Nimgaonkar VL (2015) Large-scale generation of human iPSC-derived neural stem cells/early neural progenitor cells and their neuronal differentiation Organogenesis 10(4): 365-77. |
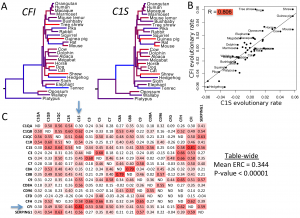 | Priedigkeit N, Wolfe N, Clark NL (2015) Evolutionary Signatures amongst Disease Genes Permit Novel Methods for Gene Prioritization and Construction of Informative Gene-Based Networks PLoS Genet 11(2):e1004967. |
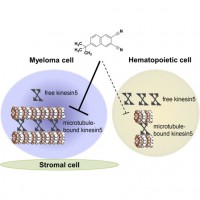 | Chattopadhyay S, Stewart AL, Mukherjee S, Huang C, Hartwell KA, Miller PG, Subramanian R, Carmody LC, Yusuf RZ, Sykes DB, Paulk J, Vetere A, Vallet S, Santo L, Cirstea DD, Hideshima T, Dančík V, Majireck MM, Hussain MM, Singh S, Quiroz R, Iaconelli J, Karmacharya R, Tolliday NJ, Clemons PA, Moore MA, Stern AM, Shamji AF, Ebert BL, Golub TR, Raje NS, Scadden DT, Schreiber SL (2015) Niche-Based Screening in Multiple Myeloma Identifies a Kinesin-5 Inhibitor with Improved Selectivity over Hematopoietic Progenitors Cell Rep. 10(5): 755-70. |
| Godin SK, Meslin C, Kabbinavar F, Bratton-Palmer DS, Hornack C, Mihalevic MJ, Yoshida K, Sullivan M, Clark NL, Bernstein KA (2015) Evolutionary and Functional Analysis of the Invariant SWIM Domain in the Conserved Shu2/SWS1 Protein Family from Saccharomyces cerevisiae to Homo sapiens Genetics 199(4): 1023-33. | |
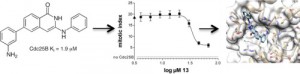 | George Rosenker KM, Paquette WD, Johnston PA, Sharlow ER, Vogt A, Bakan A, Lazo JS, Wipf P (2015) Synthesis and biological evaluation of 3-aminoisoquinolin-1(2H)-one based inhibitors of the dual-specificity phosphatase Cdc25B Bioorg Med Chem 23(12): 2810-8. |
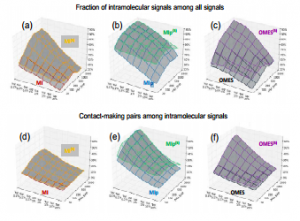 | Mao W, Kaya C, Dutta A, Horovitz A, Bahar I (2015) Comparative Study of the Effectiveness and Limitations of Current Methods for Detecting Sequence Coevolution Bioinformatics 31(12): 1929-37. |
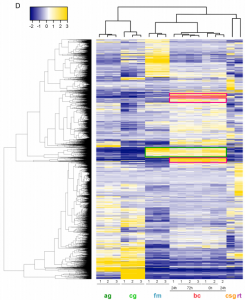 | Digestive Organ in the Female Reproductive Tract Borrows Genes from Multiple Organ Systems to Adopt Critical Functions (2015) Meslin C, Plakke MS, Deutsch AB, Small BS, Morehouse NI, Clark NL Mol Biol Evol 32(6): 1567-80. |
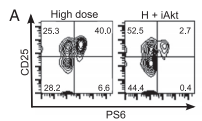 | Hawse WF, Sheehan RP, Miskov-Zivanov N, Menk AV, Kane LP, Faeder JR, Morel PA (2015) Cutting Edge: Differential Regulation of PTEN by TCR, Akt, and FoxO1 Controls CD4+ T Cell Fate Decisions J Immunol 194(10): 4615-9. |
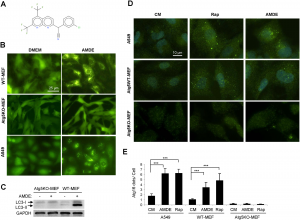 | Li M, Yang Z, Vollmer LL, Gao Y, Fu Y, Liu C, Chen X, Liu P, Vogt A, Yin XM (2015) AMDE-1 Is a Dual Function Chemical for Autophagy Activation and Inhibition PLoS One 10(3):e0122083. |
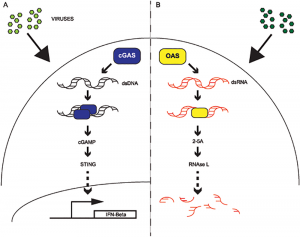 | Hancks DC, Hartley MK, Hagan C, Clark NL, Elde NC (2015) Overlapping Patterns of Rapid Evolution in the Nucleic Acid Sensors cGAS and OAS1 Suggest a Common Mechanism of Pathogen Antagonism and Escape PLoS Genet 11(5):e1005203. |
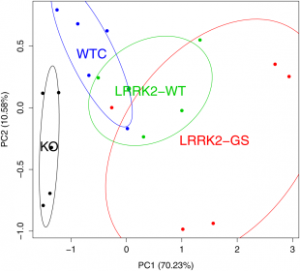 | Chikina MD, Gerald CP, Li X, Ge Y, Pincas H, Nair VD, Wong AK, Krishnan A, Troyanskaya OG, Raymond D, Saunders-Pullman R, Bressman SB, Yue Z, Sealfon SC (2015) Low-Variance RNAs Identify Parkinson's Disease Molecular Signature in Blood Mov Disord. (6):813-21.. |
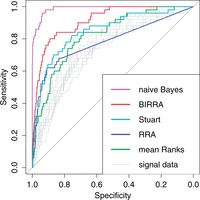 | Badgeley MA, Sealfon SC, Chikina MD (2015) Hybrid Bayesian-rank integration approach improves the predictive power of genomic dataset aggregation Bioinformatics 31(2):209-15. |
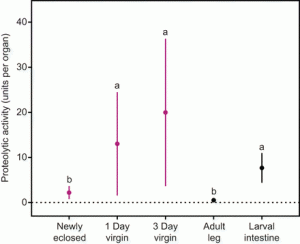 | Plakke MS, Deutsch AB, Meslin C, Clark NL, Morehouse NI (2015) Dynamic digestive physiology of a female reproductive organ in a polyandrous butterfly J Exp Biol. 18(Pt 10):1548-55. |
| Senutovitch N, Vernetti L, Boltz R, DeBiasio R, Gough A, Taylor DL (2015) Fluorescent protein biosensors applied to microphysiological systems Exp Biol Med (Maywood) 240(6): 795-808. | |
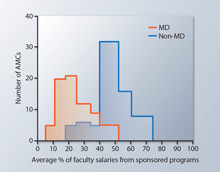 | Levine AS, Alpern RJ, Andrews NC, Antman K, Balser JR, Berg JM, Davis PB, Fitz JG, Golden RN, Goldman L, Jameson JL, Lee VS, Polonsky KS, Rappley MD, Reece EA, Rothman PB, Schwinn DA, Shapiro LJ, Spiegel AM (2015) Research in academic medical centers: Two threats to sustainable support Sci Transl Med. 7(289):289fs22. |
 | Travers T, Shao H, Joughin BA, Lauffenburger DA, Wells A, Camacho CJ (2015) Tandem phosphorylation within an intrinsically disordered region regulates ACTN4 function Sci Signal 8(378):ra51. |
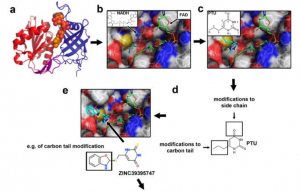 | Rahaman MM, Reinders FG, Koes D, Nguyen AT, Mutchler SM, Sparacino-Watkins C, Alvarez RA, Miller MP, Cheng D, Chen BB, Jackson EK, Camacho CJ, Straub AC (2015) Structure Guided Chemical Modifications of Propylthiouracil Reveal Novel Small Molecule Inhibitors of Cytochrome b5 Reductase 3 that Increase Nitric Oxide Bioavailability J Biol Chem 290(27):16861-72. |
| Slack RL, Spiriti J, Ahn J, Parniak MA, Zuckerman DM, Ishima R (2015) Structural integrity of the ribonuclease H domain in HIV-1 reverse transcriptase Proteins 290(27):16861-72. | |
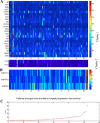 | Villaruz LC, Huang G, Romkes M, Kirkwood JM, Buch SC, Nukui T, Flaherty KT, Lee SJ, Wilson MA, Nathanson KL, Benos PV, Tawbi HA (2015) MicroRNA expression profiling predicts clinical outcome of carboplatin/paclitaxel-based therapy in metastatic melanoma treated on the ECOG-ACRIN trial E2603 Clin Epigenetics 7(1):58. |
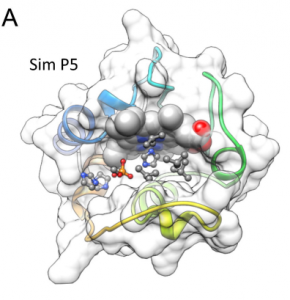 | Bakan A, Kapralov AA, Bayir H, Hu F, Kagan VE, Bahar I (2015) Inhibition of Peroxidase Activity of Cytochrome c: De Novo Compound Discovery and Validation Mol Pharmacol 88(3):421-7. |
| Koes DR, Camacho CJ (2015) Indexing Volumetric Shapes with Matching and Packing Knowl Inf Syst. 43(1):157-180. | |
| Cheng MH, Block E, Hu F, Cobanoglu MC, Sorkin A, Bahar I (2015) Insights into the Modulation of Dopamine Transporter Function by Amphetamine, Orphenadrine, and Cocaine Binding Front Neurol. 6:134. | |
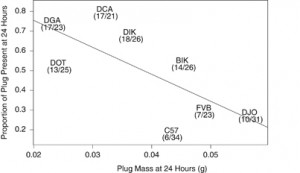 | Mangels R, Young B, Keeble S, Ardekani R, Meslin C, Ferreira Z, Clark NL, Good JM, Dean MD (2015) Genetic and phenotypic influences on copulatory plug survival in mice Heredity (Edinb) 115, 496–502. |
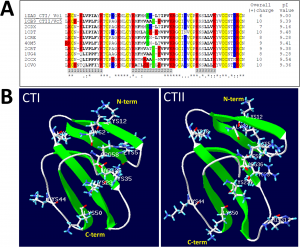 | Gasanov SE, Shrivastava IH, Israilov FS, Kim AA, Rylova KA, Zhang B, Dagda RK (2015) Naja naja oxiana Cobra Venom Cytotoxins CTI and CTII Disrupt Mitochondrial Membrane Integrity: Implications for Basic Three-Fingered Cytotoxins PLoS One 10(6):e0129248. |
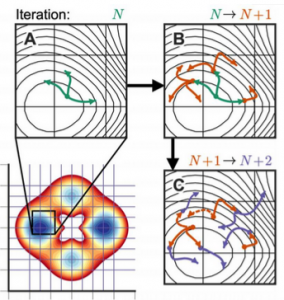 | Suárez E, Pratt AJ, Chong LT, Zuckerman DM (2015) Estimating First Passage Time Distributions from Weighted Ensemble Simulations and non-Markovian Analyses Protein Sci. 25: 67–78. |
| Cooper GF, Bahar I, Becich MJ, Benos PV, Berg J, Espino JU, Glymour C, Jacobson RC, Kienholz M, Lee AV, Lu X, Scheines R; Center for Causal Discovery team (2015) The Center for causal discovery of biomedical knowledge from Big Data J Am Med Inform Assoc. 22(6):1132-6. | |
 | Bahar I, Cheng MH, Lee JY, Kaya C, Zhang S (2015) Structure-Encoded Global Motions and Their Role in Mediating Protein-Substrate Interactions Biophys J. 109 (6): p1101–1109. |
 | Chylek LA, Harris LA, Faeder JR, Hlavacek WS (2015) Modeling for (physical) biologists: an introduction to the rule-based approach Phys Biol. 12(4): 045007. |
 | Pickett CL, Corb BW, Matthews CR, Sundquist WI, Berg JM (2015). Toward a sustainable biomedical research enterprise: Finding consensus and implementing recommendations Proc Natl Acad Sci U S A. [Epub ahead of print]. |
| Vernetti LA, Senutovitch N, Boltz R, DeBiasio R, Ying Shun T, Gough A, Taylor DL (2015). A human liver microphysiology platform for investigating physiology, drug safety, and disease models Exp Biol Med (Maywood). 241(1):101-14. | |
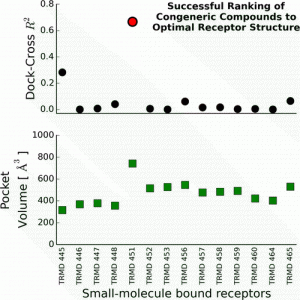 | Baumgartner MP, Camacho CJ (2015). On Choosing the Optimal Rigid Receptor for Docking and Scoring in the CSAR 2013/2014 Experiment J Chem Inf Model. [Epub ahead of print]. |
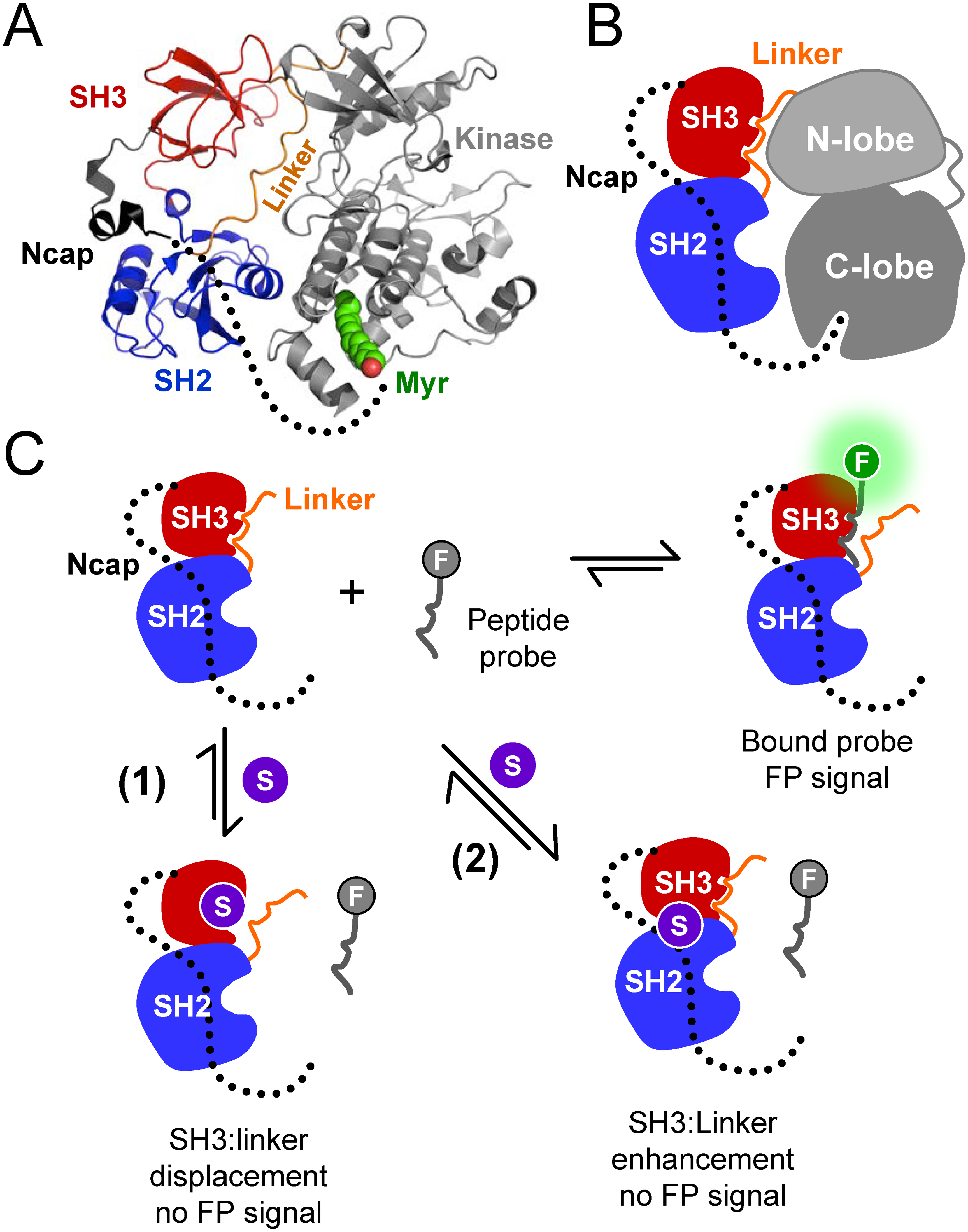 | Grover P, Shi H, Baumgartner M, Camacho CJ, Smithgall TE (2015). Fluorescence Polarization Screening Assays for Small Molecule Allosteric Modulators of ABL Kinase Function PLoSOne. 10(7):e0133590. |
 | Wolfe NW, Clark NL (2015) ERC Analysis: web-based inference of gene function via Evolutionary Rate Covariation Bioinformatics 31(23):3835-7. |
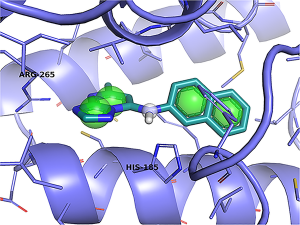 | Koes DR, Pabon NA, Deng X, Phillips MA, Camacho CJ (2015) A Teach-Discover-Treat Application of ZincPharmer: An Online Interactive Pharmacophore Modeling and Virtual Screening Tool PLoS One 10(8):e0134697. |
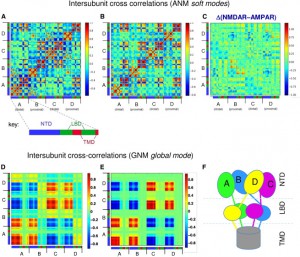 | Dutta A, Krieger J, Lee JY, Garcia-Nafria J, Greger IH, Bahar I (2015) Cooperative Dynamics of Intact AMPA and NMDA Glutamate Receptors: Similarities and Subfamily-Specific Differences Structure S0969-2126(15)00282-8. |
 | Krieger J, Bahar I, Greger IH (2015) Structure, Dynamics, and Allosteric Potential of Ionotropic Glutamate Receptor N-Terminal Domains Biophys J. S0006-3495(15)00673-6. |
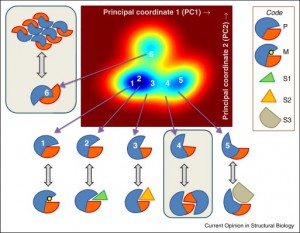 | Haliloglu T, Bahar I (2015) Adaptability of protein structures to enable functional interactions and evolutionary implications Curr Opin Struct Biol Volume 35: 17–23. |
| Chandran UR, Luthra S, Santana-Santos L, Mao P, Kim SH, Minata M, Li J, Benos PV, DeWang M, Hu B, Cheng SY, Nakano I, Sobol RW (2015) Genom Data 5:333-336. | |
| Hartmann BM, Thakar J, Albrecht RA, Avey S, Zaslavsky E, Marjanovic N, Chikina M, Fribourg M, Hayot F, Schmolke M, Meng H, Wetmur J, García-Sastre A, Kleinstein SH, Sealfon SC (2015) Human dendritic cell response signatures distinguish 1918, pandemic and seasonal H1N1 influenza viruses J Virol [Epub ahead of print]. | |
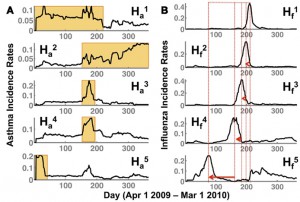 | Ramanathan A, Pullum LL, Hobson TC, Stahl CG, Steed CA, Quinn SP, Chennubhotla CS, Valkova S (2015) Discovering Multi-Scale Co-Occurrence Patterns of Asthma and Influenza with Oak Ridge Bio-Surveillance Toolkit Front Public Health 3:182. |
| Abdelraheem EM, Camacho CJ, Dömling A (2015) Focusing on shared subpockets - new developments in fragment-based drug discovery Expert Opin Drug Discov 10(11):1179-87. | |
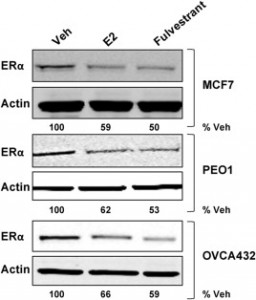 | Boisen MM, Andersen CL, Sreekumar S, Stern AM, Oesterreich S (2015) Treating Gynecologic Malignancies with Selective Estrogen Receptor Downregulators (SERDs): Promise and Challenges Mol Cell Endocrinol. 418 Pt 3:322-33. |
| Keskin O, Dyson HJ, Bahar I (2015) Biomolecular Systems Interactions, Dynamics, and Allostery: Reflections and New Directions Biophys J. 109(6):E01-2. | |
 | Zwier MC, Adelman JL, Kaus JW, Pratt AJ, Wong KF, Rego NB, Suárez E, Lettieri S, Wang DW, Grabe M, Zuckerman DM, Chong LT (2015) WESTPA: An interoperable, highly scalable software package for weighted ensemble simulation and analysis J Chem Theory Comput 11(2):800-809. |
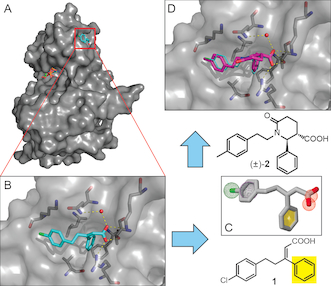 | Kroon E, Schulze JO, Süß E, Camacho CJ, Biondi RM, Dömling A (2015) Discovery of a Potent Allosteric Kinase Modulator by Combining Computational and Synthetic Methods Angew Chem Int Ed Engl. 54: 13933–13936. |
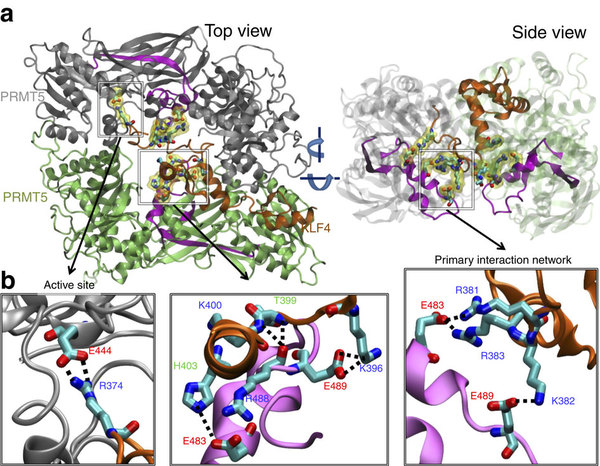 | Hu D, Gur M, Zhou Z, Gamper A, Hung MC, Fujita N, Lan L, Bahar I, Wan Y (2015) Interplay between arginine methylation and ubiquitylation regulates KLF4-mediated genome stability and carcinogenesis Nat Commun 6:8419. |
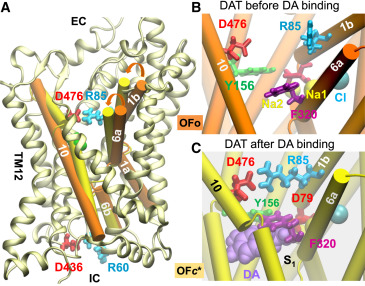 | Cheng MH, Bahar I (2015) Molecular Mechanism of Dopamine Transport by Human Dopamine Transporter Structure. 23 (11): 2171–2181. |
| Gough A, Shun T, Lansing Taylor D, Schurdak M. (2015) A Metric and Workflow for Quality Control in the Analysis of Heterogeneity in Phenotypic Profiles and Screens Methods S1046-2023(15)30126-2. | |
| Wang P, Bahreini A, Gyanchandani R, Lucas P, Hartmaier RJ, Watters RJ, Jonnalagadda AR, Trejo Bittar HE, Berg A, Hamilton RL, Kurland BF,Weiss K, Mathew A, Leone JP, Davidson NE, Nikiforova MN, Brufsky AM, Ambros TF, Stern AM, Puhalla S, Lee AW, Oesterreich S.(2015) Sensitive detection of mono- and polyclonal ESR1 mutations in primary tumors, metastatic lesions and cell free DNA of breast cancer patients Clin Cancer Res. [Epub ahead of print]. | |
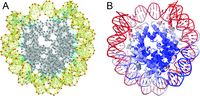 | Li H, Chang YY, Yang LW, Bahar I (2015) iGNM 2.0: the Gaussian network model database for biomolecular structural dynamics Nucleic Acids Res 44(D1):D415-22. |
| Wang S, Faeder JR, Setlow P, Li YQ (2015) Memory of Germinant Stimuli in Bacterial Spores MBio 24;6(6). | |
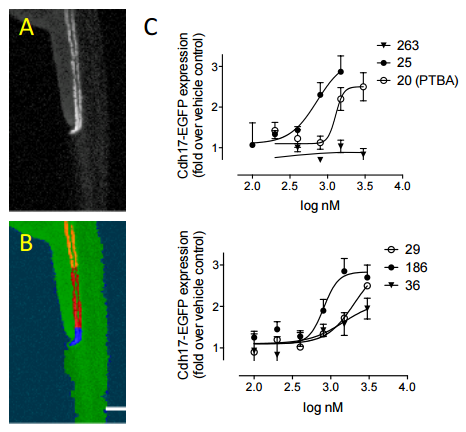 | Skrypnyk NI, Sanker S, Brilli-Skvarca L, Novitskaya T, Woods C, Chiba T, Patel K, Goldberg ND, McDermott L, Vinson PN, Calcutt MW, Huryn DM, Vernetti LA, Vogt A, Hukriede N, de Caestecker MP (2015) Delayed treatment with PTBA analogs reduces post injury renal fibrosis after kidney injury Am J Physiol Renal Physiol. [Epub ahead of print] |
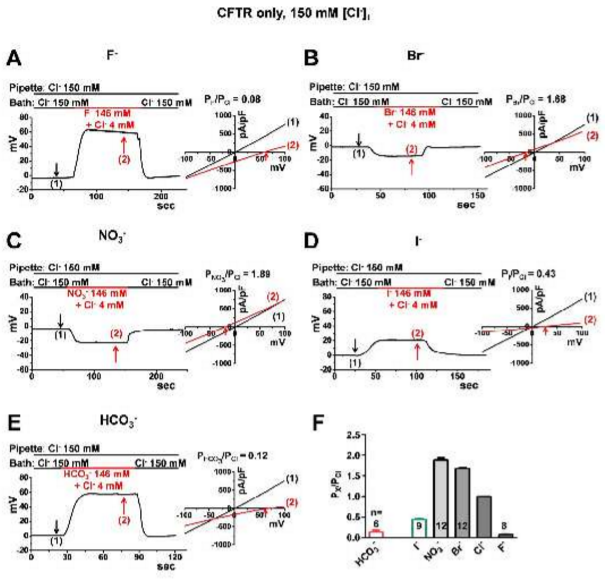 | Jun I, Cheng MH, Sim E, Jung J, Suh BL, Kim Y, Son H, Park K, Kim CH, Yoon JH, Whitcomb DC, Bahar I, Lee MG (2015) Pore dilation increases the bicarbonate permeability of CFTR, ANO1, and glycine receptor anion channels J Physiol. [Epub ahead of print] |
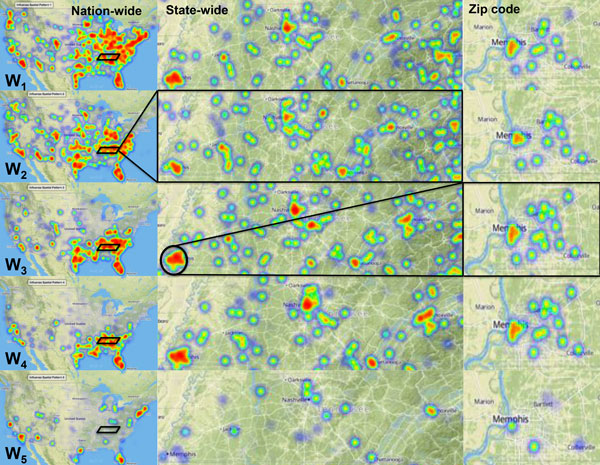 | Ramanathan A, Pullum LL, Hobson TC, Steed CA, Quinn SP, Chennubhotla CS, Valkova S (2015) ORBiT: Oak Ridge biosurveillance toolkit for public health dynamics BMC Bioinformatics 16 Suppl 17:S4. |
| Hlavacek WS, Gnanakaran S, Munsky B, Wall ME, Faeder JR, Jiang Y, Nemenman I, Resnekov O (2015) The eighth q-bio conference: meeting report and special issue preface Phys Biol. 12(6):060401. | |
 | Spiriti J, Zuckerman DM (2015) Tabulation as a high-resolution alternative to coarse-graining protein interactions: Initial application to virus capsid subunits J. Chem. Phys. 143(24):243159. |
| Gur M, Zomot E, Cheng MH, Bahar I (2015) Energy landscape of LeuT from molecular simulations J Chem Phys 143(24):243134. | |
| Savol AJ, Chennubhotla CS (2015) Approximating frustration scores in complex networks via perturbed Laplacian spectra Phys Rev E Stat Nonlin Soft Matter Phys. 92(6-1):062806. |
Other Years
‣ 2021 Publications
‣ 2020 Publications
‣ 2019 Publications
‣ 2018 Publications
‣ 2017 Publications
‣ 2016 Publications
‣ 2015 Publications
‣ 2014 Publications
‣ 2013 Publications
‣ 2012 Publications
‣ 2011 Publications
‣ 2010 Publications
‣ 2009 Publications
‣ 2008 Publications
‣ 2007 Publications
‣ 2006 Publications
‣ 2005 Publications
