“Crosstalk Between Toll-like Receptor Pathways”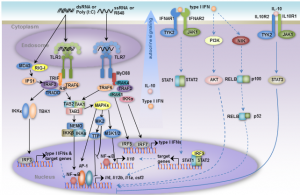
A collaboration between the Bahar lab, the Ding lab (National Singapore University), and the Thiagarajan lab (Harvard), led by Dr. Liu in the Bahar lab.
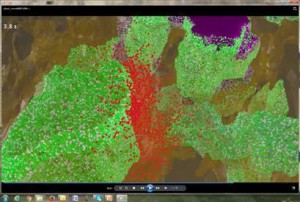
Toll-like receptors (TLRs) recognize pathogens and stimulate the innate immune response. Liu et al uncovers how the crosstalk between TLR pathways enables macrophages to fine-tune their responses to multiple infection events. In particular, macropahges can “memorize” the first pathogen attack and boost the immune response to subsequent attacks. The crosstalk also helps to avoid the harmful excessive inflammatory responses.